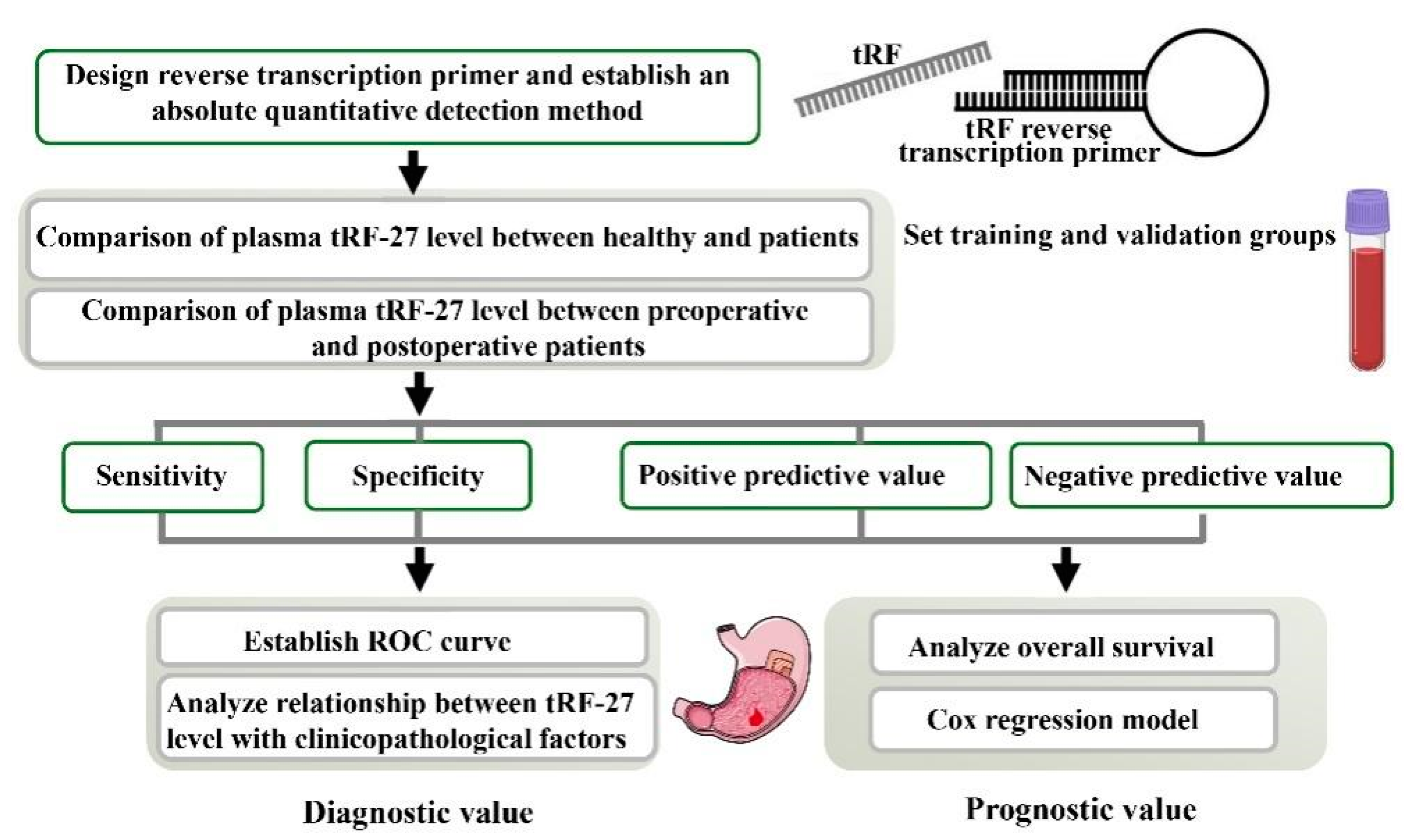
IJMS | Free Full-Text | Establishment of an Absolute Quantitative Method to Detect a Plasma tRNA-Derived Fragment and Its Application in the Non-Invasive Diagnosis of Gastric Cancer
Absolute quantification of transcription factors during cellular differentiation using multiplexed targeted proteomics
Identifying the combinatorial control of signal-dependent transcription factors | PLOS Computational Biology

Absolute quantification of transcription factors reveals principles of gene regulation in erythropoiesis | bioRxiv
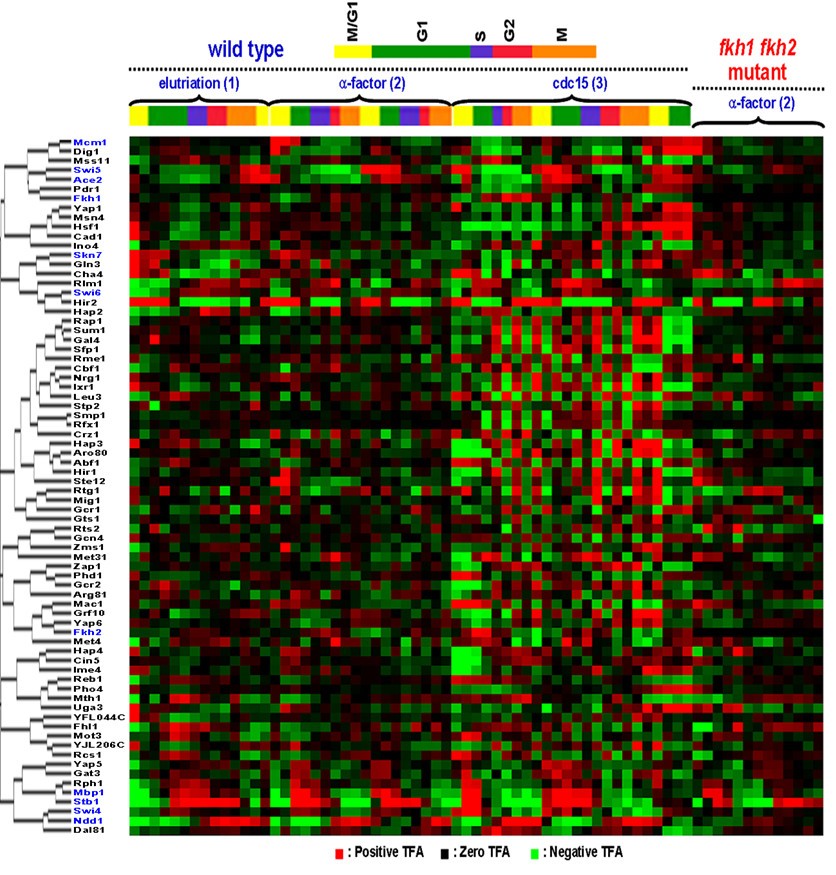
Inferring yeast cell cycle regulators and interactions using transcription factor activities | BMC Genomics | Full Text
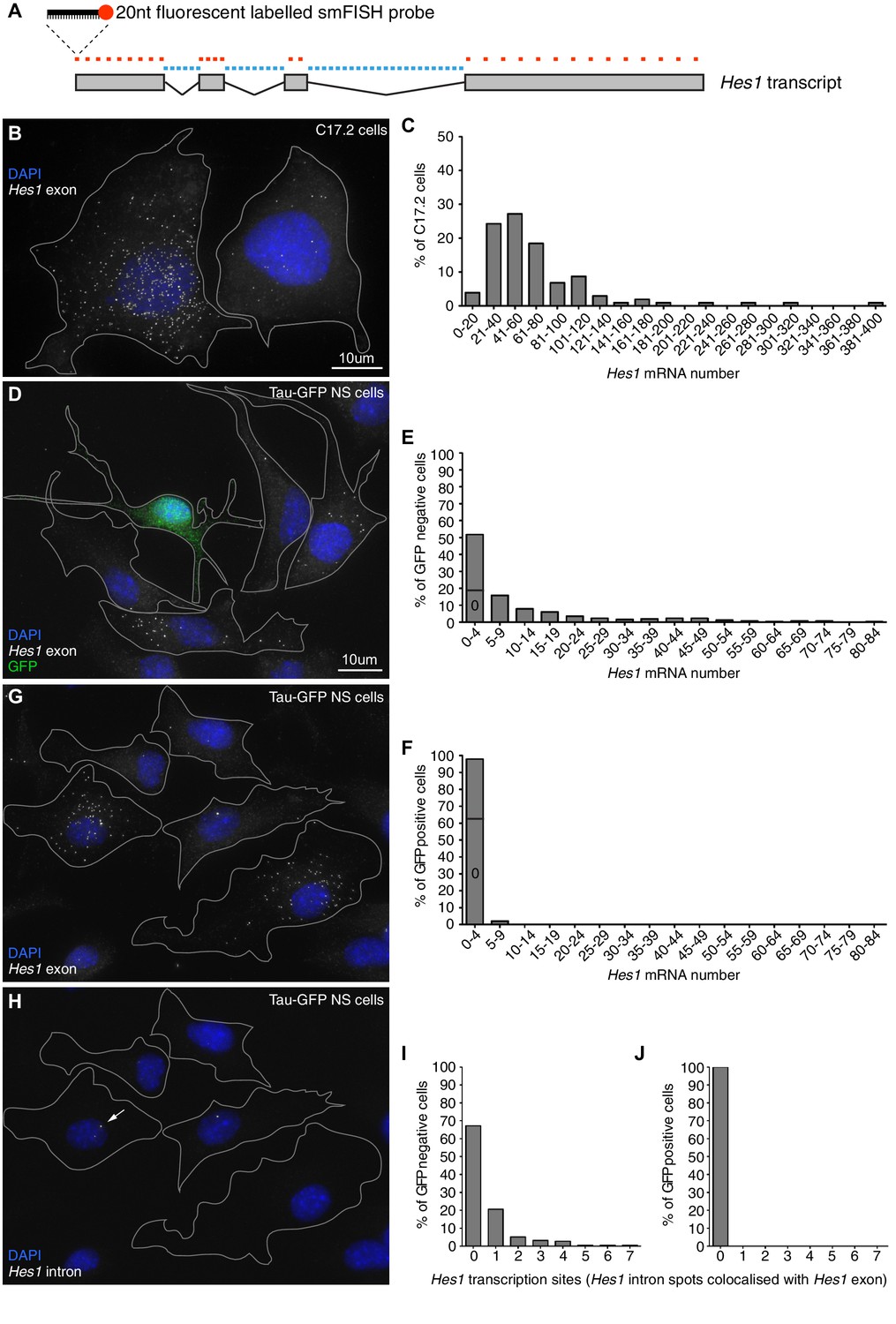
Figures and data in Stochasticity in the miR-9/Hes1 oscillatory network can account for clonal heterogeneity in the timing of differentiation | eLife
![PDF] ChIP-Seq of transcription factors predicts absolute and differential gene expression in embryonic stem cells | Semantic Scholar PDF] ChIP-Seq of transcription factors predicts absolute and differential gene expression in embryonic stem cells | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/3bfd424415e33b55adccb8434638ea9853cf997f/2-Figure1-1.png)
PDF] ChIP-Seq of transcription factors predicts absolute and differential gene expression in embryonic stem cells | Semantic Scholar